Methods & Materials
Animals
All procedures were approved by the Institutional Animal Care and were carried out in accordance with the Animal Welfare Act. Sixty male Sprague–Dawley rats were obtained from Harlan (Indianapolis, IN), housed in an environmentally controlled facility on a 12/12-h light/dark cycle, and provided chow and water ad libitum. Male Sprague-Dawley rats were randomly assigned to CON (CON, n=20), doxorubicin (DOX, n=27), or creatine + doxorubicin (CR+DOX, n= 20) groups. CR+DOX received rodent chow supplemented with 3% creatine monohydrate and the CON and DOX received standard rodent chow. After two weeks of feeding, the CR+DOX and DOX groups received DOX injection and CON received a saline injection as a placebo. Five days following injections, each animal was anesthetized using heparinized sodium pentobarbital, and when a tail pinch reflex was absent, the heart was removed and the right soleus (SOL) and right extensor digitorum longus (EDL) was excised, weight, frozen in liquid nitrogen, and stored at -80°C for future biochemical analysis.
All procedures were approved by the Institutional Animal Care and were carried out in accordance with the Animal Welfare Act. Sixty male Sprague–Dawley rats were obtained from Harlan (Indianapolis, IN), housed in an environmentally controlled facility on a 12/12-h light/dark cycle, and provided chow and water ad libitum. Male Sprague-Dawley rats were randomly assigned to CON (CON, n=20), doxorubicin (DOX, n=27), or creatine + doxorubicin (CR+DOX, n= 20) groups. CR+DOX received rodent chow supplemented with 3% creatine monohydrate and the CON and DOX received standard rodent chow. After two weeks of feeding, the CR+DOX and DOX groups received DOX injection and CON received a saline injection as a placebo. Five days following injections, each animal was anesthetized using heparinized sodium pentobarbital, and when a tail pinch reflex was absent, the heart was removed and the right soleus (SOL) and right extensor digitorum longus (EDL) was excised, weight, frozen in liquid nitrogen, and stored at -80°C for future biochemical analysis.
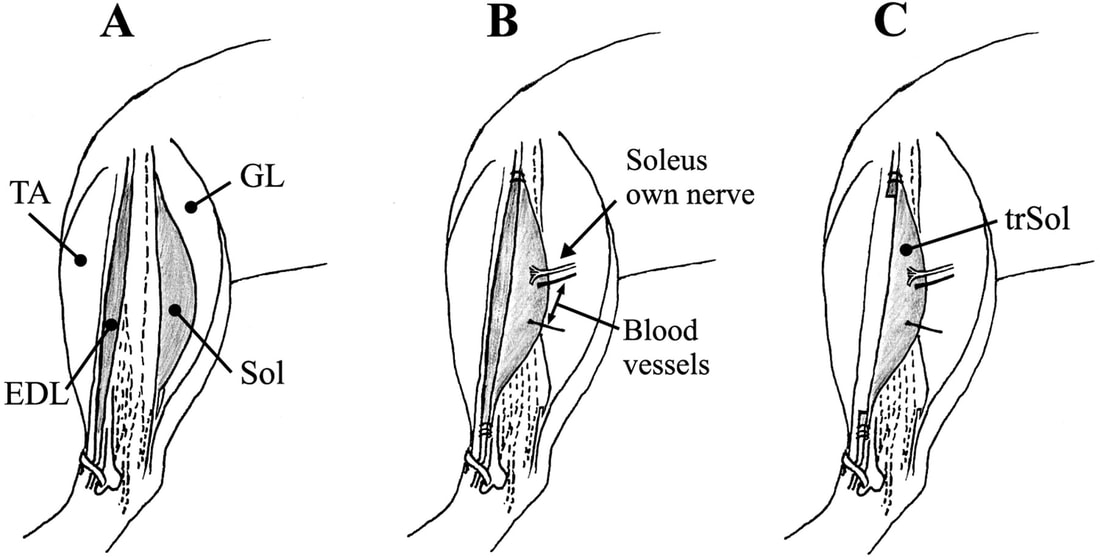
A) Muscles labeled: EDL and Sol are of interest B) When Removing Sol: only cut nerve to remove C) Entire Sol removed and used
Homogenate Preparation
The flash frozen left hindlimb muscles were homogenized in radioimmunoprecipitation assay (RIPA) buffer and protease enzyme inhibitor. After manual homogenization, samples were sonicated to increase nuclear protein recovery. Homogenates were then be spun in a microcentrifuge for 10 minutes at 3000 g at 4°C. Total protein was quantified in samples using a Genesys 20 photospectrometer. A determined amount of RIPA buffer was added to standardize protein concentration.
The flash frozen left hindlimb muscles were homogenized in radioimmunoprecipitation assay (RIPA) buffer and protease enzyme inhibitor. After manual homogenization, samples were sonicated to increase nuclear protein recovery. Homogenates were then be spun in a microcentrifuge for 10 minutes at 3000 g at 4°C. Total protein was quantified in samples using a Genesys 20 photospectrometer. A determined amount of RIPA buffer was added to standardize protein concentration.
Mortar and Pestle used to manually Centrifuge used to spin the substance to separate Photospectrometer used to quantify and
homogenize proteins from the unwanted tissue standardize proteins
homogenize proteins from the unwanted tissue standardize proteins
Western Blotting
Western blot analysis was conducted on muscle samples for the presence of MRFs in the SOL and EDL via SDS-PAGE. An equal volume of Lammeli buffer was added to samples in a microcentrifuge polypropylene vial, reducing protein concentration in half. Samples were loaded onto 4-20% gradient Tris-Glycine NuPage precast gels (LifeTechnologies: Carlsbad, CA). Gels were run at 125 V (constant voltage) and 4 mA current for 2 hours in a Xcell II blot module (Invitrogen, LifeTechnologies), until proteins had migrated the whole gel. Proteins were transferred to 0.45 micron polyvinylidene fluoride (PVDF) membranes over 90 minutes at 25 V and 100 mA. Membranes were blocked for 1 hour in 15 mL of TBST + 5% milk, and then incubated with gentle agitation overnight in 10 mL of primary antibodies. Membranes were then be washed in TBST three times for 5 minutes, followed by incubation in appropriate species-specific secondary antibodies for 1 hour. After three more 5-minute washes in TBST, membranes were prepared for protein band detection.
Detection was executed by enhanced chemiluminescence (ECL) (C-Digit, Li-Cor: Lincoln, NE), and ImagJ software (NIH: Bethesda, MD) was used to quantify protein expression. The primary antibodies of interest included rabbit monoclonal MyoD and myogenin (Santa Cruz Biotechnology: Dallas, TX). The rabbit monoclonal anti- GAPDH (Abcam) was used as a loading control. Molecular weights of protein bands were ensured in reference to a MagicMarkTM XP standard ladder (Novex, LifeTechnologies). Secondary antibodies (Santa Cruz Biotechnology) corresponded to associated species (rabbit) and include horseradish peroxidase (HRP) for adequate reactivity.
Western blot analysis was conducted on muscle samples for the presence of MRFs in the SOL and EDL via SDS-PAGE. An equal volume of Lammeli buffer was added to samples in a microcentrifuge polypropylene vial, reducing protein concentration in half. Samples were loaded onto 4-20% gradient Tris-Glycine NuPage precast gels (LifeTechnologies: Carlsbad, CA). Gels were run at 125 V (constant voltage) and 4 mA current for 2 hours in a Xcell II blot module (Invitrogen, LifeTechnologies), until proteins had migrated the whole gel. Proteins were transferred to 0.45 micron polyvinylidene fluoride (PVDF) membranes over 90 minutes at 25 V and 100 mA. Membranes were blocked for 1 hour in 15 mL of TBST + 5% milk, and then incubated with gentle agitation overnight in 10 mL of primary antibodies. Membranes were then be washed in TBST three times for 5 minutes, followed by incubation in appropriate species-specific secondary antibodies for 1 hour. After three more 5-minute washes in TBST, membranes were prepared for protein band detection.
Detection was executed by enhanced chemiluminescence (ECL) (C-Digit, Li-Cor: Lincoln, NE), and ImagJ software (NIH: Bethesda, MD) was used to quantify protein expression. The primary antibodies of interest included rabbit monoclonal MyoD and myogenin (Santa Cruz Biotechnology: Dallas, TX). The rabbit monoclonal anti- GAPDH (Abcam) was used as a loading control. Molecular weights of protein bands were ensured in reference to a MagicMarkTM XP standard ladder (Novex, LifeTechnologies). Secondary antibodies (Santa Cruz Biotechnology) corresponded to associated species (rabbit) and include horseradish peroxidase (HRP) for adequate reactivity.